Disordered Structure Refinement (DSR)
Enhanced modelling and refinement of disordered structures with SHELXL
Preface
The following user manual explains the usage of the program DSR. It allows for a semi-automatic modelling of disordered moieties. Its database contains molecular fragments and their corresponding restraints as well as a fitting procedure to place these fragments on the desired position in the unit cell.
The web page can be found at https://dkratzert.de/dsr.html and the development platform at https://github.com/dkratzert/DSR.
If you find any bugs in this program, have feature requests or just comments; please don’t hesitate to write an email to <dkratzert@gmx.de> to report these.
Please cite DSR as:
D. Kratzert, J.J. Holstein, I. Krossing, J. Appl. Cryst. 2015 48, 933-938. doi:10.1107/S1600576715005580
Disclaimer
You are responsible for the correctness of restraints applied to your crystal structure. The restraints applied by DSR are only suggestions to stabilize a first model. DSR does not replace the judgment of an expert.
Program Overview
The program-package consists of a simple text-database with fragments of molecules and the DSR program itself. It acts as a preprocessor for SHELXL .res files. The user has to insert a special command line in the SHELXL .res file and the DSR program reads this information. The command lines main purpose is to tell DSR, how to orient a molecular fragment from the database in the unit cell. In practice, the user has to choose a minimum of three target positions (atoms or Q-peaks) in the structure and the corresponding atoms from the database fragment (source atoms) that should be placed on the target positions.
You prefer Olex2? Then you might want to use the FragmentDB plugin. It is a port of DSR to Olex2.
Installation
Windows
Execute the “DSR-setup-[version number].exe” and follow the instructions. DSR needs at least Windows 7 or Windows 10 for the latest version. If you install DSR the first time, a restart of Windows might be necessary.
Linux
Either install DSR according to the installation procedure of your LINUX distribution. Or install the dsr-shelx package from pypi.org. DSR expects a shelxl or xl executable version 2013 or above in the system path.
The update from DSR for python 2.7 (version 236 and below) to DSR for Python 3 needs some manual work. You need to uninstall the old version and make sure the files dsr.sh, dsr-mac and dsr-linux are removed from /etc/profile.d/ and/or /etc/paths.d/.
Do the following steps in order to install and run DSR: Some Linux distributions like Ubuntu need a python3-venv package to be installed first.
>> cd
>> python3 -m venv dsrenv <-- creates a virtual environment
>> source dsrenv/bin/activate # (in Windows: dsrenv\Scripts\activate.bat) <-- Activates the environment
>> pip install dsr-shelx
>> sudo ln -s /home/[username]/dsrenv/bin/dsr /usr/local/bin/dsr <-- link the executable to a global accessible directory
>> dsr
MacOS
Install the dsr-shelx package from pypi.org.
Easiest way in macOS might be:
Install homebrew: https://brew.sh/
brew install pipx
pipx install dsr-shelx
Create a link that is usable for all users:
sudo ln -s /Users/[username]/.local/bin/dsr /usr/local/bin/dsr
The update from DSR for python 2.7 (version 236 and below) to DSR for Python 3 needs some manual work. You need to uninstall the old version and make sure the files dsr.sh, dsr-mac and dsr-linux are removed from /etc/profile.d/ and/or /etc/paths.d/.
User-defined database
DSR expects the database file “dsr_user_db.txt” with self-made fragments in the user’s home directory. The installation procedures create an empty user database in the following directories if no previous database exists:
Windows: C:\users\username\
Linux: /home/username/
Mac OS X: /Users/username/
DSR in ShelXle
Since version 181, ShelXle has the ability to start a graphical user interface for DSR. A mouse click on Tools–> DSR plugin (figure 1) will start the DSR GUI (figure 2):
Figure 1
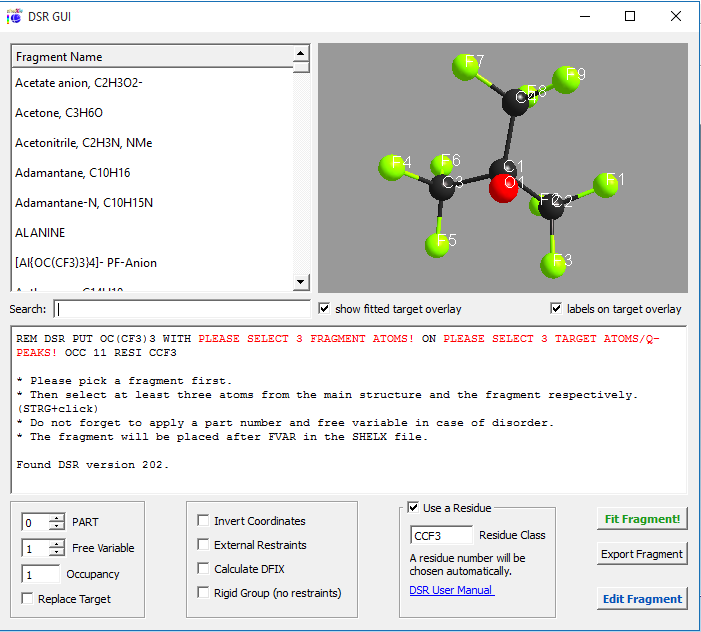
Figure 2
Now you need to select a fragment in the list. The list of fragments can be searched using the search field. The search shortens the list to the fragments that best match by name.
To fit a fragment into the structure in ShelXle, select three atoms/Q-peaks in the target molecule (with a left mouse click while holding STRG) and the fragments 3D view (just left mouse click) each. The 3D view should now show a preview of the fitted fragment (figure 3):
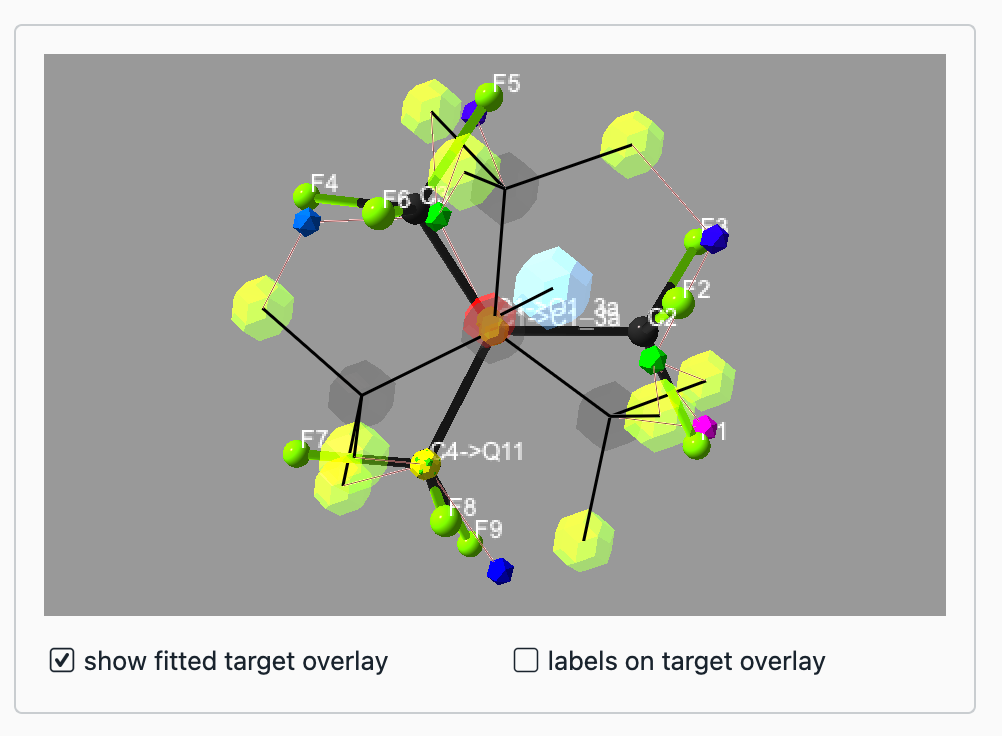
Figure 3
You can now control all the features of DSR with the options menu below (figure 4):
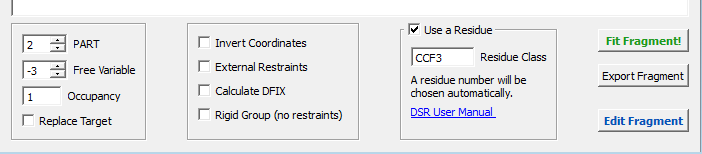
Figure 4
Setting PART to zero will disable them. The residue number will always be chosen as the next free available. You can safely leave this as it is or change the residue name.
The “Free variable” option defines the free variable for the fragment occupation in SHELXL. The Free variable will be combined with the occupation option. For example a free variable of –3 and an occupation of 1 will be combined to –31. The result appears instantly in the output window.
“External restraints” writes the restraints to an external file.
“Calculate DFIX” automatically generates DFIX/DANG/FLAT restraints from the geometry of the fragment. This can be particular useful to stabilize the a fragment on special positions.
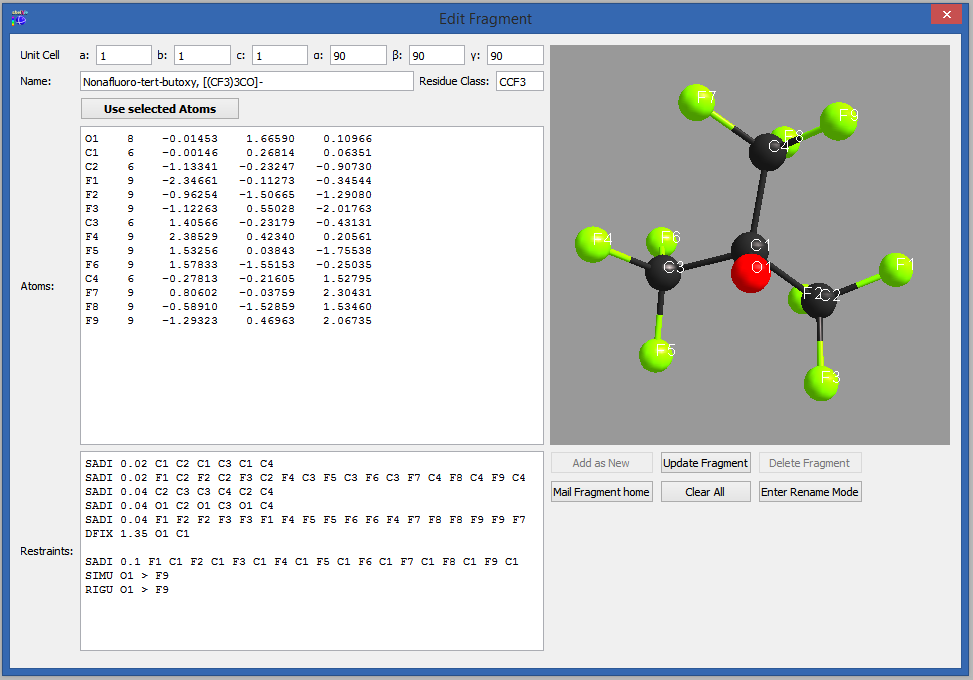
Figure 5
To create or edit a fragment, click on "Edit fragment". The edit window allows adding, updating and deleting of fragments.
Similar to the syntax in "dsr_usr_db.txt", you can choose to define the atom type by the name of the atom or with a negative atomic number.
They will be stored in the users fragment database "dsr_usr_db.txt" in your home directory. Different to the fragment creation by hand, you do not have to invent a database name tag. It will be randomly chosen, because the GUI will never show them. Instead the GUI always shows the real fragments names.
Rename mode
In order to rename the atoms in a fragment, click on “Enter Rename Mode”. The editor will you now only allow editing of atomic names. The restraints will be renamed accordingly while you are typing. Accept the changes with “OK” or discard them with “Abort”.
After renaming, you can save the changes by clicking “Add as new” or “Update fragment”.
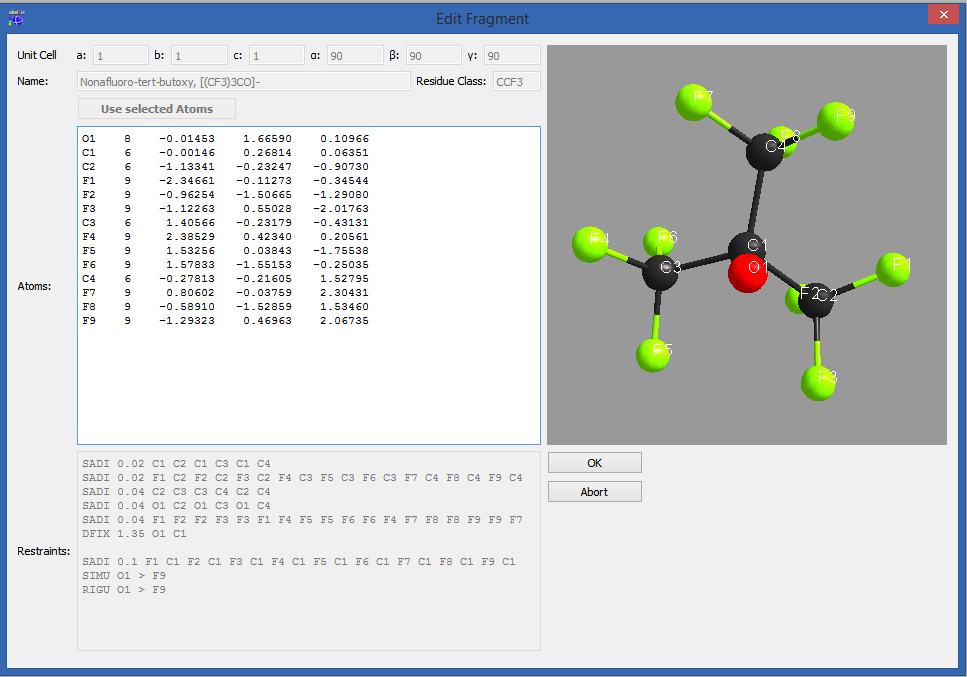
Figure 6
Background Informtion
General Procedure
Although DSR has a graphical user interface in ShelXle, it can also be run on the command line only. Therefore, insert the DSR command explained in the following chapter into the .res file. Then, run “dsr -r filename.res” and DSR will transfer the fragment from the database into the structure. The resulting filename.res can now be reopened for further refinement. The new fragment is then exactly in front of the HKLF instruction of the .res file. The respective restraints are located directly after the UNIT instruction. In order to be able to revert the changes in the .res file, DSR creates a backup file in a subdirectory "dsrsaves" with the current date as file name before every action.
Command Syntax
The DSR command has the following syntax:
REM DSR PUT/REPLACE fragment WITH atom1 atom2 atom3 ... ON atom2 atom3
atom4 ... PART n OCC mn RESI class num [alias] DFIX
The command is introduced with a REM because SHELXL should never interpret the DSR command line.
PUT Put the fragment on there, ignoring atoms on this position.
REPLACE Replace the target atoms. Hydrogen atoms of target atoms should be prior removed.
fragment The name of the desired molecule or fragment.
WITH Behind WITH are the source atoms. They are at least three atoms from the fragment.
ON Behind ON are the target atoms. They are at least three atoms or
Q-peaks in the .res- file.
[atom n] Minimum three atoms each (including Q-peaks). Source and
target have to include the same number of atoms and/or Q-peaks. Target
atoms can be either regular atoms or atoms in residues. Atoms in
residues can be addressed by the “_” notation. C1_2 would be atom C1 in
residue number 2.
PART n Optional SHELXL PART definition.
OCC mn Optional occupancy and free variable definition for the fragment.
DFIX Optional, generates DFIX/DANG restraints instead of those from the
database. All 1,2- and 1,3-distances in the fragment are restrained with
DFIX and DANG respectively. DSR also searches for rings in the fragment
and generates FLAT restraints for flat rings.
RESI class num [alias] Optional residue definition as in SHELXL.
SPLIT Only for a disordered CF3 group on two positions (CF6 fragment).
Splits the pivot atom in two positions.
Example
The following command line can be inserted anywhere between the atoms of a .res file.
REM DSR put toluene with C1 C2 C3 on Q1 C5 C2
The command is always introduced with a REM. The case does not matter, DSR is completely case insensitive. The DSR command line can be up to two lines with a trailing "=" for a continuation line like in SHELX. Please note that the second line of the DSR COMMAND after the "=" must begin with a leading whitespace.
The minimal requirement for DSR to work is rem dsr put/replace “fragment” with “three atoms/Q-peaks” on “three atoms/Q-peaks”.
The new molecule or fragment is placed just before the HKLF instruction. DSR applies a new naming scheme to the fragment while inserting it into the .res file. Essentially it searches if any atom name from the database fragment is already used in the .res file. If this applies, the program places a suffix letter (A, B, ...) to the atom name in the .res file. This renaming is completely turned off if residues are used. Atoms of the new fragment are then addressed by their residue.
putDSR uses the coordinates of the given atoms/Q-peaks and puts the fragment on these coordinates leaving the given atoms in place. The above example will place the fragment on the coordinates of Q1, C5 and C2. The atoms C5 and C2 would remain where they were located before.
replaceDSR searches for the coordinates of the given atoms/Q-peaks but in contrast to the former example, it replaces the target atoms and all atoms in 1.3 Å distance around each atom of the fitted fragment that are in PART 0. This mode is useful to quickly rename atoms from a solution by SHELXT.
It is highly advised to use residues with DSR. They make many things easier and DSR takes care about all details regarding residues. Normally it is sufficient to simply use the RESI command without any options in DSR. This way, DSR takes the residue class from the database and finds the next residue number automatically. Restraints for the same residue class are only introduced once. Also the atoms in the fragment would not be renamed:
REM DSR put toluene with C1 C2 C3 on Q1 C5 C2 RESI
Use of Residues
To use the RESI command in DSR has several advantages. It places the fragment into a residue and therefore no renaming of the atoms in the fragment needs to be performed by DSR. If residues are used, the restraints like "SADI_class Atoms" are inserted only once, since they act on the atoms in all residues with the same class together.
In SHELXL you define residues as:
RESI 1 abc
atoms ...
RESI 0
RESI 2 abc
atoms ...
RESI 0
RESI 3 xyz
atoms ...
RESI 0
A big plus is that you can use restraints like "SAME_class C1 > C20". This command will apply a SAME restraint to all residues in class. A single atom inside a residue is now addressed with Atom_number, like "DFIX 1.5 C1_2 C3_0". Generally, residue 0 includes all atoms outside of residues.
Residues are especially useful if the same moiety is repeated several times in a crystal structure. And that is what DSR is intended for! Different moieties of the same residue class are distinguished by different residue numbers. A residue number must be unique in a .res file. The DSR command RESI without any further options is usually the best practice. DSR then uses the residue class name from the database and finds the next free residue number by itself. But the user can also specify a particular residue class and/or number after the RESI command, if desired.
Please be aware that SIMU behaves special with residues. Let’s assume we have a disordered phenyl group on two positions where the two rings are slightly rotated along their bond to the next part of the molecule (Figure 7):
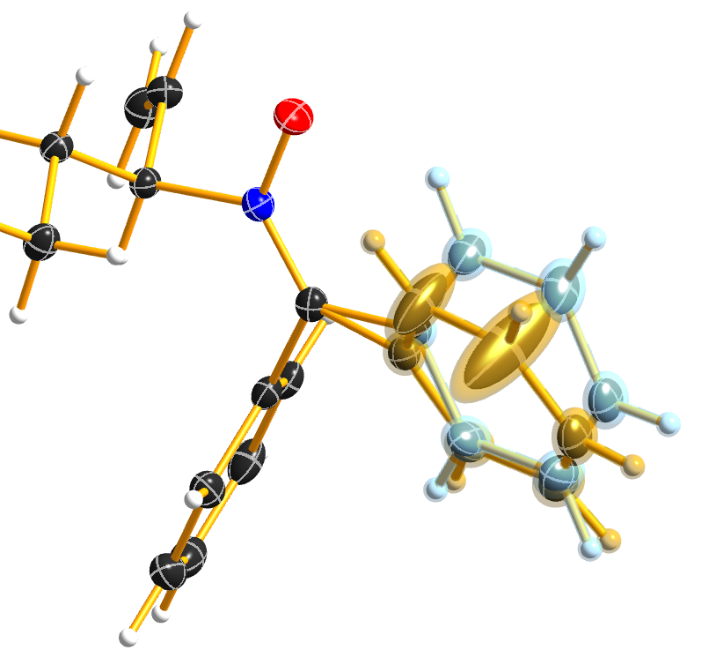
Figure 7
DSR introduces, among others, the restraint “SIMU_BENZ C1 > C6”. This is not wrong, but still not enough in every case, because SHELXL only generates SIMU restraints inside each residue. In this case, it generates “SIMU C1_1 C2_1 C3_1 C4_1 C5_1 C6_1” and “SIMU C1_2 C2_2 C3_2 C4_2 C5_2 C6_2”. But because of the close proximity and the small rotational movement we can assume that the thermal parameter in both disorder parts are more equal. Therefore, we have to include all involved atoms explicitly in one SIMU command: “SIMU C1_1 > C6_1 C1_2 > C6_2”. This can possibly be optimized by more than one SIMU and different values for the standard deviation and dmax, e.g. “SIMU 0.02 0.04 0.5 atoms” and “SIMU 0.04 0.08 1.3 atoms” (Figure 8).
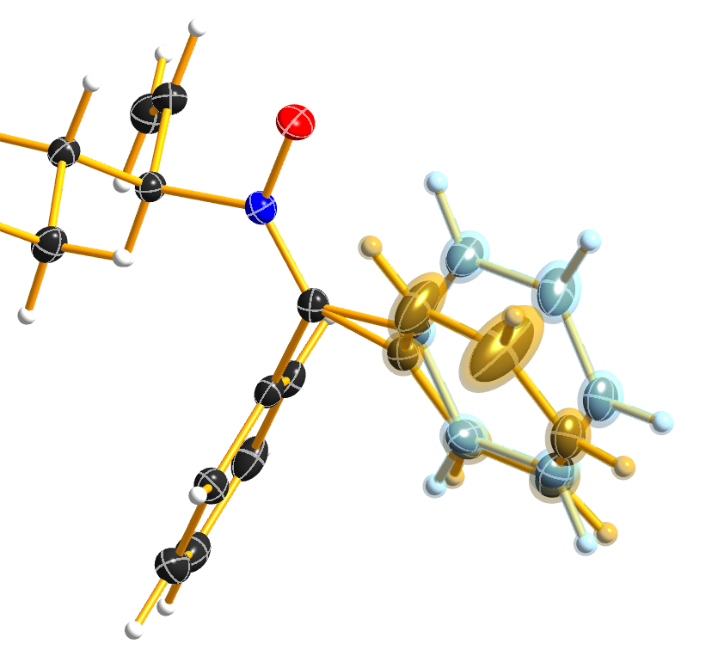
Figure 8
The RESI option of DSR can be used in three ways:
If only a RESI command is given (best practice), the residue class is taken from the database entry and the residue number is automatically generated.
If RESI with only a number is given, DSR takes the residue class from the database with the given number.
RESI with a number and a class overwrites the information from the database and gives complete control over the residue.
A given class, number or alias always overwrites the information of the database.
The manual on the SHELX website gives more detailed information about residues: https://shelx.uni-goettingen.de/wikis.php
Common Problems with residues
When using residues you may encounter the following warning from SHELXL:
** No match for C1A C2A in SIMU **
These errors are most likely due to a different arrangement of atoms within the same residue class or to a different number of atoms within the same class.
Residues of same class always have to have the same arrangement of atoms and the same number of atoms!
For example
SIMU_foo C1 > C3 RESI 1 foo C1 1 ... C2 1 ... C3 1 ... RESI 2 foo PART 1 21 C1 1 ... C2 1 ... C3 1 ... PART 2 C1A 1 ... C2A 1 ... C3A 1 ... PART 0 RESI 0
would produce the above error, because RESI 2 has six atoms and RESI 1 only three.
You can get rid of the error if you move the PART 2 out of the residue. Therefore, move RESI 0 before PART 2:
SIMU_foo C1 > C3 RESI 1 foo C1 1 ... C2 1 ... C3 1 ... RESI 2 foo PART 1 21 C1 1 ... C2 1 ... C3 1 ... RESI 0 PART 2 C1A 1 ... C2A 1 ... C3A 1 ... PART 0
Here, RESI 1 and RESI 2 have the same number of atoms and the atoms of PART 2 are irrelevant for the residues.
Command line Syntax
Following options are available in the Windows or Unix command line to control the behavior of DSR:
usage: dsr [-h] [-r "res file"] [-re "res file"] [-e "fragment"] [-c "fragment"] [-t] [-i "tgz file"] [-l] [-n]
optional arguments:
–h, --help Show a help message and exit.
–r "res file" res file with DSR command. Usually this option is used to process the SHELXL file with DSR.
–re "res file" Same as "-r", but a file called dsr_class_name.dfx or dsr_class_number_name.dfx is written which includes the restraints for the fragment for the .res file "name" in the residue "class" and "number".
–e "fragment" Exports a fragment from the database to the file [fragment].res. It includes the minimal requirements to view the fragment in a 3D molecule viewer. If a PLATON executable and ImageMagic installation is in the system path, it also creates a .png-picture of the molecule.
–c "fragment" Exports the fragment to the clipboard with Cartesian coordinates. This fragment can for example be used for modelling in the program Olex2.
–t Inverts the current fragment. Available for fragment fit, import and export.
–I "GRADE file" Imports a molecular fragment from .tgz file of the Grade server http://grade.globalphasing.org/ into the dsr_usr_db.txt.
–l Displays all fragments in the database with the line numbers where they occur.
–s "string" Search the database for given string.
–g Keep the fragment as rigid group (AFIX 9). The fragment will only move as a whole. Restraints will be omitted.
–u Updates DSR to the most recent version if any available. (In Linux, you need super-user rights to perform an update)
–n Only transfers the fragment. The fragment fit after the fragment transfer is disabled.
Database Format Definition
The database format was deliberately kept very simple. It consists of a system database in the dsr_db.txt and a user database in the dsr_user_db.txt. The system database is overwritten with every new program install while the user database will always stay untouched. So the user can easily add new fragments to its own dsr_user_db.txt database. The syntax mainly follows the SHELXL syntax:
<fragment name> <- Start tag
RESI class <- Required, defines the residue name of db entry.
restraints <- Any restraints and comments following the
SHELXL syntax.
You must enter at least one restraint!
e.g. RIGU C1 > C7
FRAG 17 a b c alpha beta gamma <- FRAG card with AFIX number and cell parameters.
Atom sfac-number coordinates <- One isotropic atom per line following SHELX syntax:
O1 1 1.2345 0.6734 0.8352 <- Either the atom type is recognized by the atom name
for positive Numbers in the second column.
C1 -6 0.2683 0.4783 0.1616 <- Or the atom type is defined by the negative atomic
number in the second column.
</fragment name> <- End tag. Same as start tag but with a slash.
Anything not being an atom after FRAG is ignored.
Fragment names CF3, CF6 and CF9 are reserved by DSR. Do not attempt to use them in database entries.
Only lines beginning with valid SHELXL instructions are allowed in the header.
Anything behind the 5th column in the atom list is ignored.
Long lines can be wrapped with an equal sign (=) and a space character in the next line like in SHELXL, but the can also be of any length. All lines will be wrapped to fit in the SHELXL file automatically.
Database Example
A usual database entry looks like the following:
<Toluene>
rem CCDC: BUWME
rem Name: Toluene, C7H8
RESI TOL
SAME C2 > C6 C1
SAME C1 C6 < C2 C7
HFIX 137 C7
FLAT C1 > C7
SIMU C1 > C7
RIGU C1 > C7
FRAG 17 11.430 12.082 15.500 106.613 100.313 90.68
C1 1 0.268330 0.478380 0.161680
C2 1 0.205960 0.555770 0.217990
C3 1 0.249400 0.600760 0.310040
C4 1 0.357300 0.568990 0.348900
C5 1 0.420800 0.492470 0.294060
C6 1 0.376630 0.447580 0.201340
C7 1 0.221500 0.430400 0.060360
</TOLUENE>
The restraints applied by DSR might be stricter than necessary. After introduction of a new fragment, the refinement can be proceeded as usual. In the course of you should review the restraints. Modifications to database fragments should always be done in the dsr_user_db.txt and not in the dsr_db.txt. The user database will not be overwritten during updates. The fragment names must be unique in both databases. Every valid restraints from SHELXL can be used, even HFIX is possible.
The syntax follows the SHELXL syntax. All entries between the start tag <Toluene> and the FRAG command are considered as the database entry header. Comments can be introduced with REM. All lines with non-SHELX commands are ignored.
If a “rem Name:” statement is given, the name after this statement is printed in the list of available fragments.
After FRAG until the end tag </TOLUENE> only atoms with SHELX syntax are accepted.
Step by Step Working Example (command line)
You can find the following example in the DSR install directory. This example explains the procedures of using DSR via the command line. You can also use the graphical user interface for DSR in ShelXle if you do not like to type DSR commands (see chapter ShelXle Integration).
Step 0
Open “p21c.res”.
Residue number 3 turns out to be a part of a disorder.
Apply a PART and the free variable 2 to this residue with “PART 1 21”:
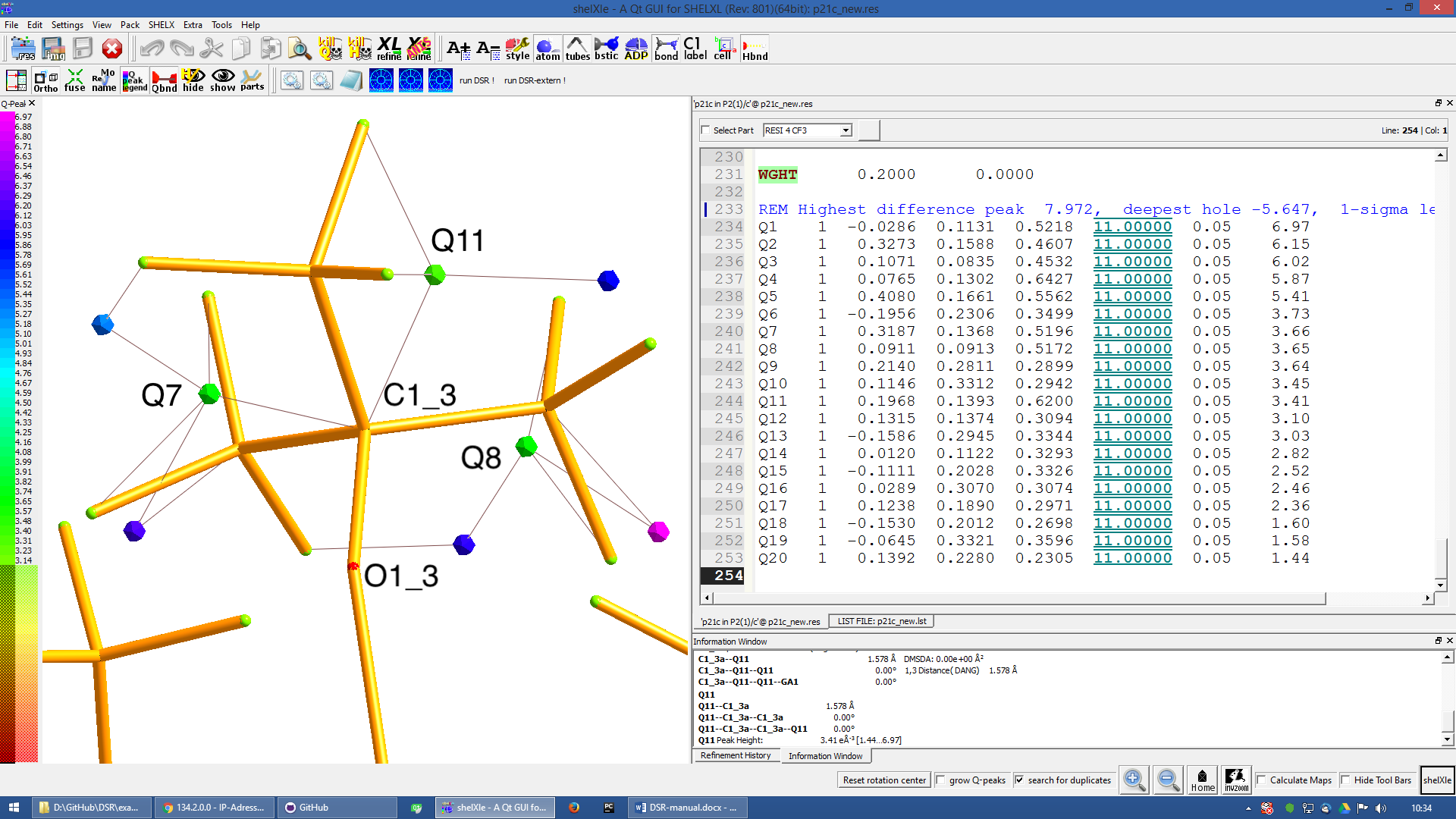
Figure 9
RESI 3 CCF3
PART 1 21
O1 3 0.156860 0.210330 0.529750 21.00000 0.05000
C1 1 0.198400 0.149690 0.543840 21.00000 0.05000
C2 1 0.283540 0.125060 0.490330 21.00000 0.05000
F1 4 0.357580 0.075160 0.511570 21.00000 0.05000
F2 4 0.359970 0.170660 0.471080 21.00000 0.05000
F3 4 0.212170 0.104000 0.438130 21.00000 0.05000
C3 1 0.086960 0.103360 0.549830 21.00000 0.05000
F4 4 0.118510 0.041260 0.547430 21.00000 0.05000
F5 4 -0.003540 0.114200 0.501240 21.00000 0.05000
F6 4 0.032040 0.112650 0.606550 21.00000 0.05000
C4 1 0.279960 0.152770 0.609930 21.00000 0.05000
F7 4 0.222220 0.188240 0.652750 21.00000 0.05000
F8 4 0.297220 0.093900 0.636590 21.00000 0.05000
F9 4 0.395450 0.177140 0.602220 21.00000 0.05000
PART 0
RESI 0
Step 1
Now you can insert the command line for DSR after the residue 3.
The command is
rem dsr put oc(cf3)3 with O1 c1 c2 on O1_3 C1_3 Q11 part 2 occ -21 resi
to place the fragment OC(CF3)3 on the position of O1_3 C1_3 q11.
In addition we want to have the fragment in a PART 2 with the occupancy of −21 and in a residue. DSR automatically finds a free residue number and uses a residue name from the database. All these options are placed in one line. As usual in SHELXL, lines longer than 80 characters can be continued with "=" at the end and a whitespace before the first character in the next line.
Save the res file after editing.
Step 2
Now run "dsr -r p21c.res" on the Windows/Unix command line.
DSR will run over the res file, insert the fragment and makes a refinement with “L.S. 0”. This finally inserts the fragment. You can see the status before the refinement in the “p21c_step2.ins” file.
D:\tmp\example>dsr -r p21c.res
--------------------------------- D S R -- v242 ----------------------------
No residue number was given. Using residue number 5.
Inserting oc(cf3)3 into res File.
Source atoms: O1, C1, C2
Target atoms: O1_3, C1_3, Q11
RESI instruction is enabled. Leaving atom numbers as they are.
Fragment atom names: O1, C1, C2, F1, F2, F3, C3, F4, F5, F6, C4, F7, F8, F9
-----------------------------------------------------------------------------
Running SHELXL with "C:\bn\sxtl\xl.exe -b3000 p21c" and "L.S. 0"
SHELXL Version 2019/3
wR2 = 0.5454
GooF = 6.4660
R1 = 0.2101
Runtime: 0.3 s
DSR run complete.
Reopen the resulting res file.
Step 3
The fragment turned out to be successfully fitted on its desired position:
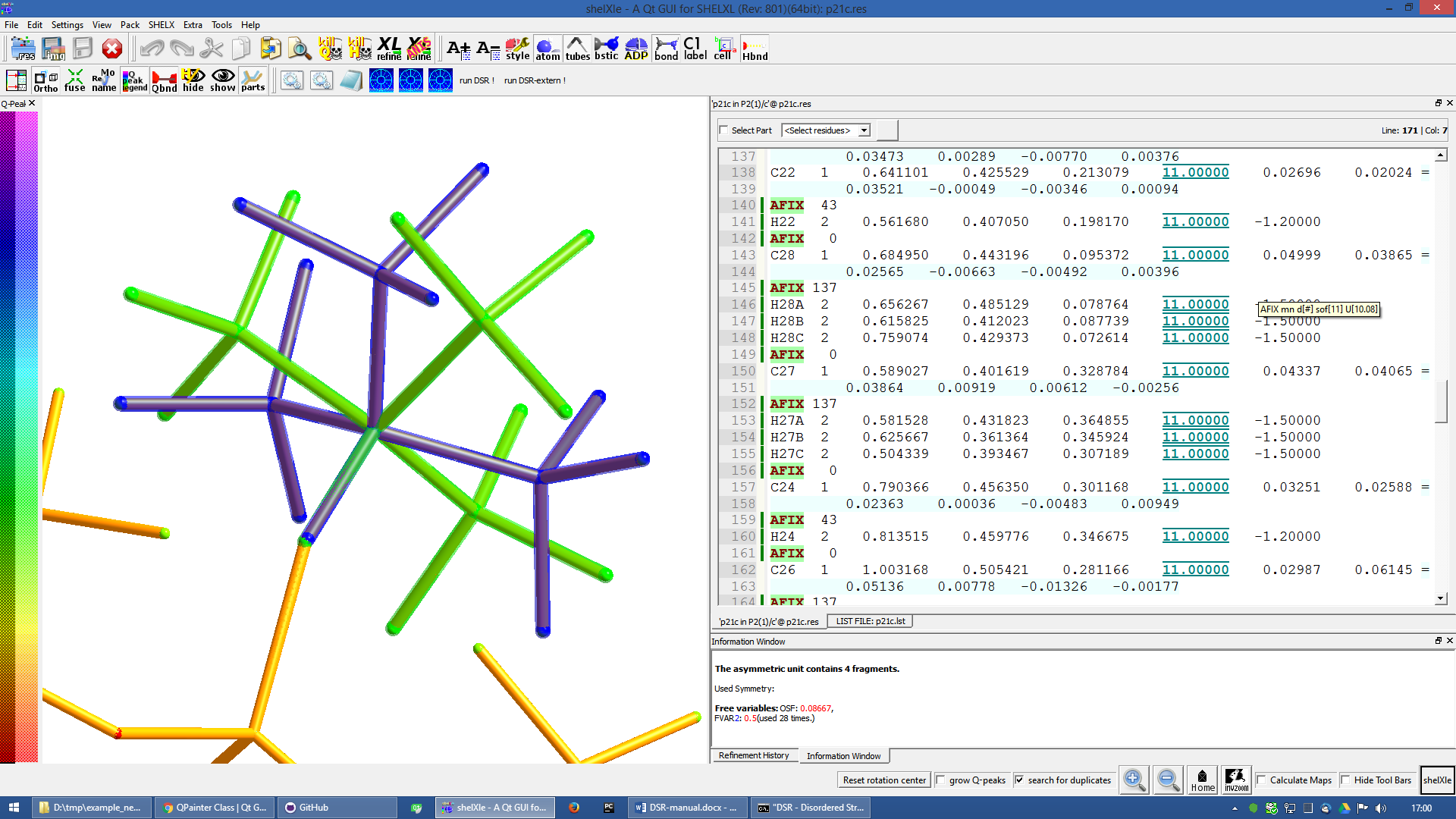
Figure 10
The previously used DSR command line is now removed and will not be recognized by DSR again.
Step 4
Do the same procedure from step 1 onwards to refine the second disorder.
The final model of the anion should look like this:
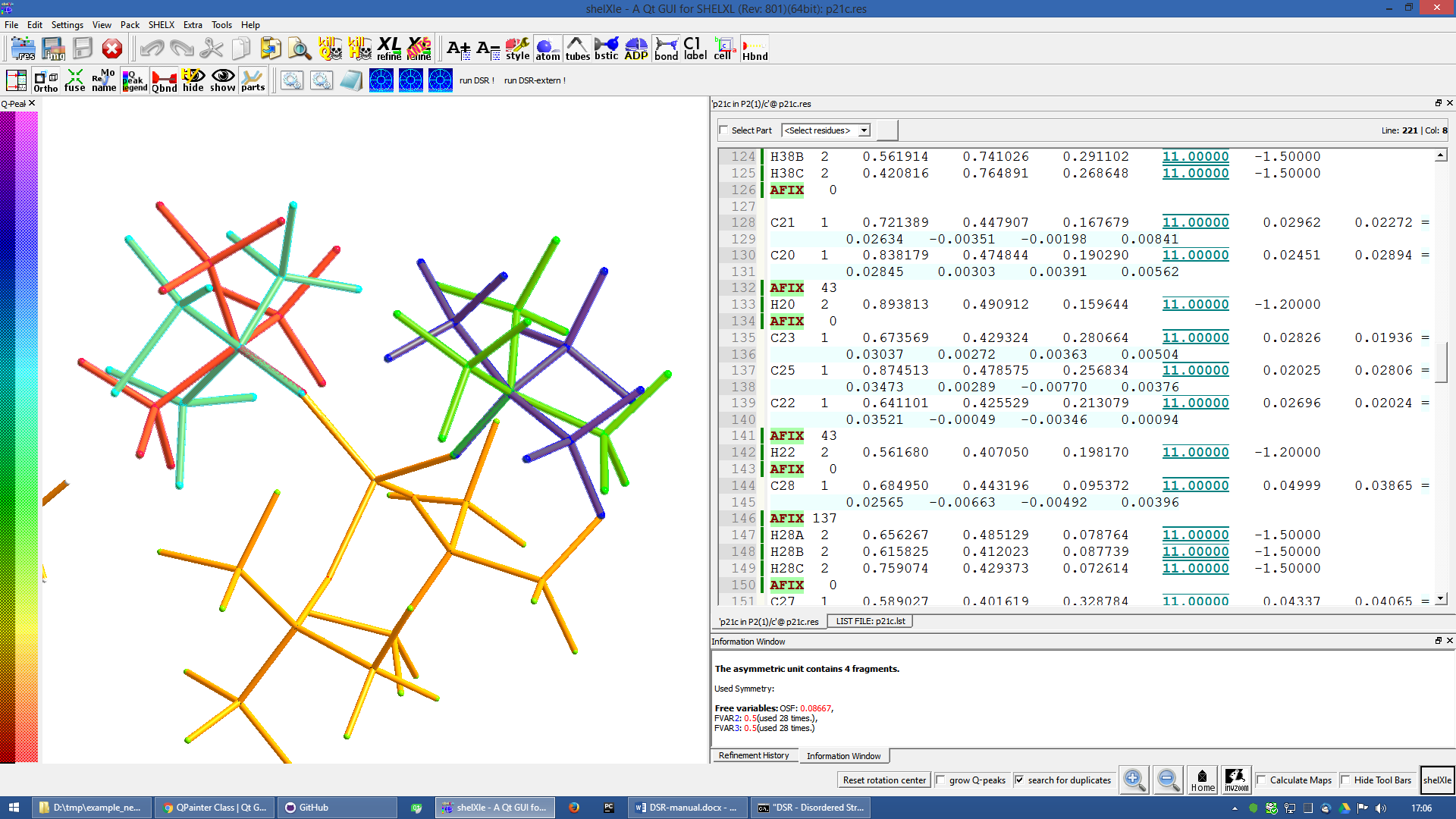
Figure 11
Now you can add/remove additional restraints and further refine the structure as usual. Already existing restraints for an existing residue class will not be inserted again, because they already act for all residues together (with SADI_CCF3 for example).
A good assumption for the model would also be that all Al–O distances are the same. Therefore, we should add the restraint
SADI Al1_0 O1_1 Al1_0 O1_2 Al1_0 O1_3 Al1_0 O1_4 Al1_0 O1_5 Al1_0 O1_6
The central oxygen and carbon atoms of the disordered perfluorinated tert-butyl alcohol group are now in close proximity. Therefore, it is a good idea to safe parameters and stabilize their ADPs with an EADP constraint:
EADP C1_3 C1_5
EADP O1_5 O1_3
EADP C1_6 C1_4
EADP O1_4 O1_6
After ten cycles of refinement, the structure should produce an R1 of 4% and should have no bigger residual density features left over (0.30 eÅ:sup:-3 level):
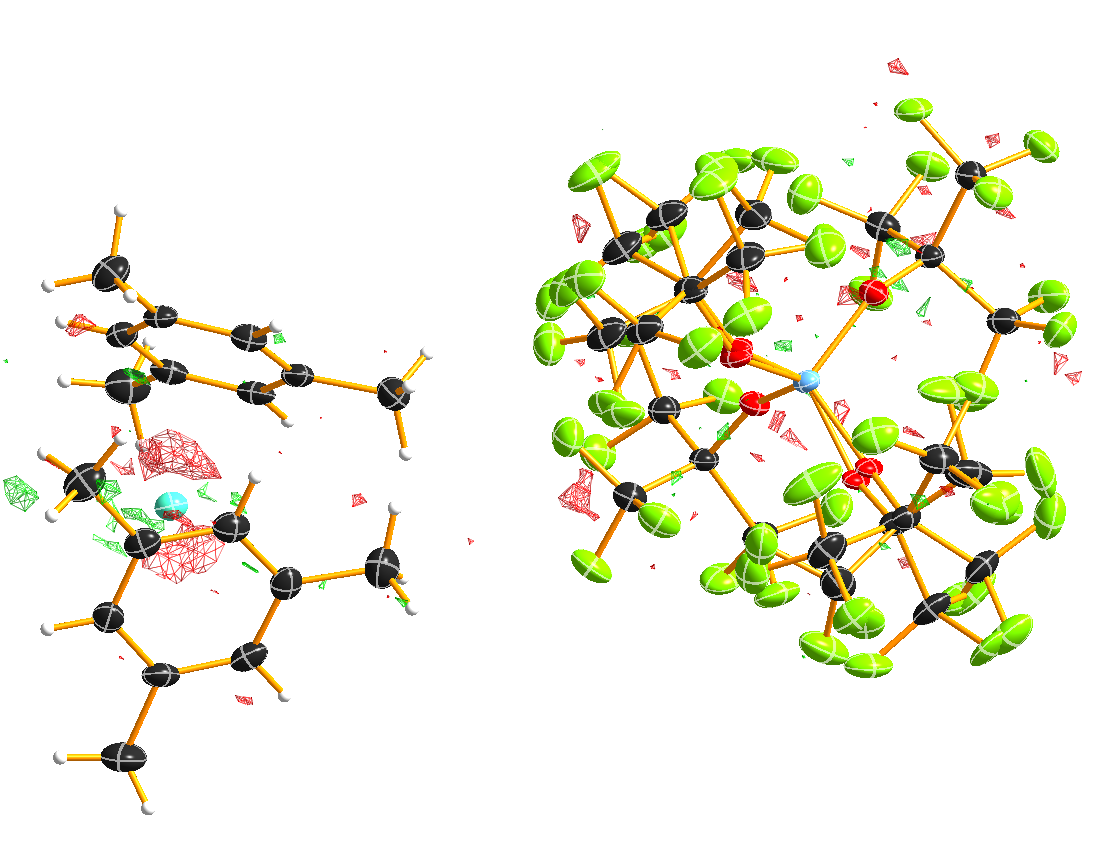
Figure 12
Import fragments from GRADE
GRADE from Global Phasing Ltd. http://grade.globalphasing.org is a ligand restraint generator whose main source of restraint information is the Cambridge Structural Database (CSD) of small-molecule crystal structures, queried using the MOGUL program developed by the CCDC. Where small-molecule information is lacking, Grade uses quantum chemical procedures to obtain the restraint values.
Fragments obtained by GRADE can be imported to DSR with the –i command line option. The GRADE server outputs a .tgz file including several files. Execution of “dsr –i filename.tgz” will import a GRADE fragment from these files. The fragment gets imported to the “dsr-user-db.txt” in the DSR program directory. You also might need to change the fragment and residue name after the import. The best way is to supply the GRADE server with a .mol2 file. This way you can choose the atom names and their sorting yourself. mol2 files can be generated if you create a molecule with Avogadro (save as .res file) <http://sourceforge.net/projects/avogadro/> then you must rename the atoms and open the res file with mercury <http://www.ccdc.cam.ac.uk/Solutions/CSDSystem/Pages/Mercury.aspx>. Mercury can now save a .mol2 file for GRADE.
CF3-Groups
DSR is able to generate CF3 groups with the respective restraints automatically. In contrast to the other fragments, for modelling of CF3 groups with DSR you only need to define one single target atom (a carbon atom). The CF3 group can be modeled either as ordered group on one position (like an AFIX 130 would do), as well as a disordered group on two or three positions. The respective fragment names are CF3, CF6 and CF9. In ShelXle, select the respective carbon atom, and click the “fit fragment” button. On the command line, a disordered CF3 group on three positions would be modelled using:
REM DSR put CF9 on C1
The result is the following plus the respective free variables added to the FVAR line.
REM CF3 group made by DSR:
SUMP 1 0.0001 1 2 1 3 1 4
SADI 0.02 C1 F1B C1 F2B C1 F3B C1 F4B C1 F5B C1 F6B C1 F7B C1 F8B C1 F9B
SADI 0.04 F1B F2B F2B F3B F3B F1B F4B F5B F5B F6B F6B F4B F7B F8B F8B F9B =
F9B F7B
SADI 0.1 C2 F1B C2 F2B C2 F3B C2 F4B C2 F5B C2 F6B C2 F7B C2 F8B C2 F9B
RIGU C2 C1 F1B > F9B
PART 1 21
F1B 3 0.128560 -0.462510 0.547150 11.00000 0.04
F2B 3 0.047696 -0.372991 0.451178 11.00000 0.04
F3B 3 0.040064 -0.244730 0.559034 11.00000 0.04
PART 2 31
F4B 3 0.118185 -0.486145 0.500565 11.00000 0.04
F5B 3 0.020565 -0.289148 0.471484 11.00000 0.04
F6B 3 0.077571 -0.304938 0.585313 11.00000 0.04
PART 3 41
F7B 3 0.086249 -0.450791 0.462663 11.00000 0.04
F8B 3 0.017551 -0.238494 0.514080 11.00000 0.04
F9B 3 0.112520 -0.390946 0.580619 11.00000 0.04
PART 0
Existing fluorine atoms connected to the respective carbon atom will be deleted by DSR beforehand.
If you like DFIX instead of SADI, add DFIX to the DSR command line:
REM DSR put CF9 on C1 DFIX
The special command SPLIT in combination with the CF6 fragment tries to split the target carbon atom in two positions. The coordinates of the two positions are the principal axes of the carbon atom ellipsoid in its longest direction. The restraints will be adjusted accordingly.
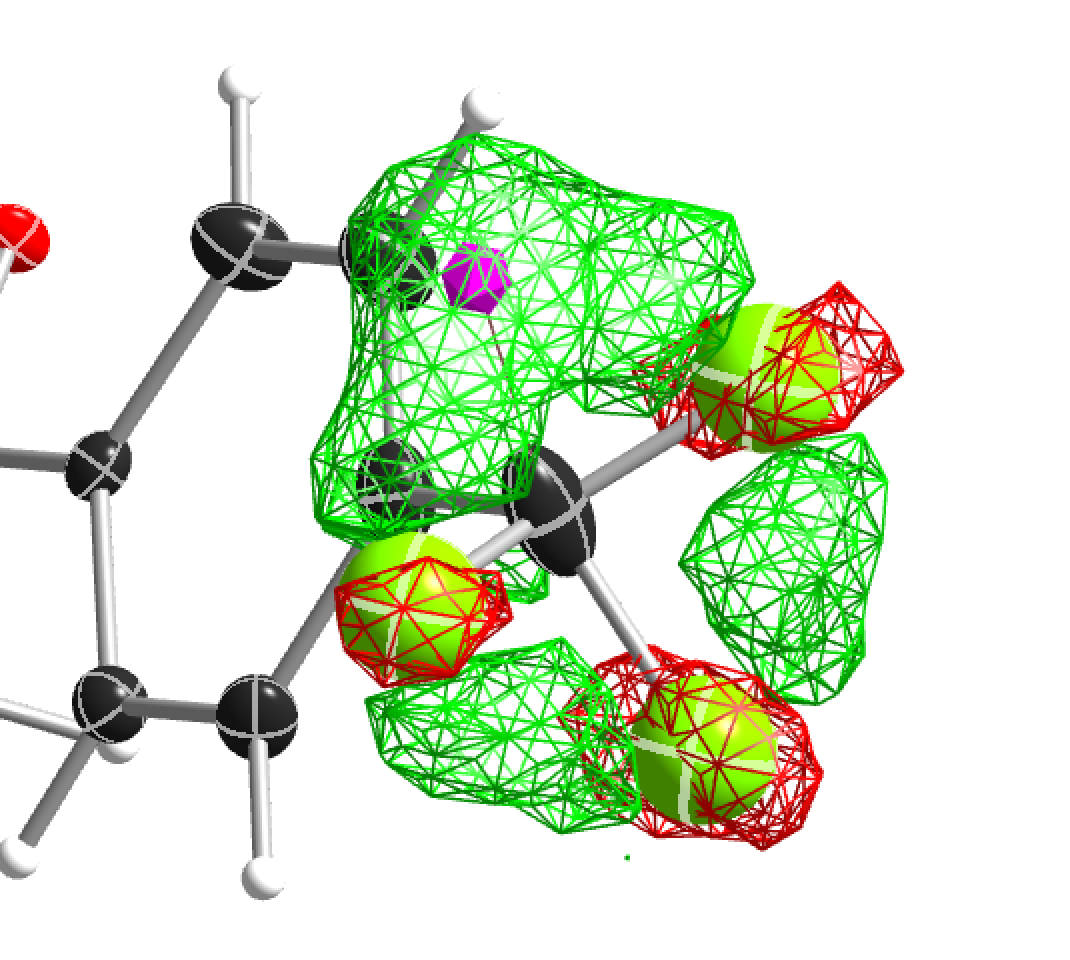
Figure 13 with C1
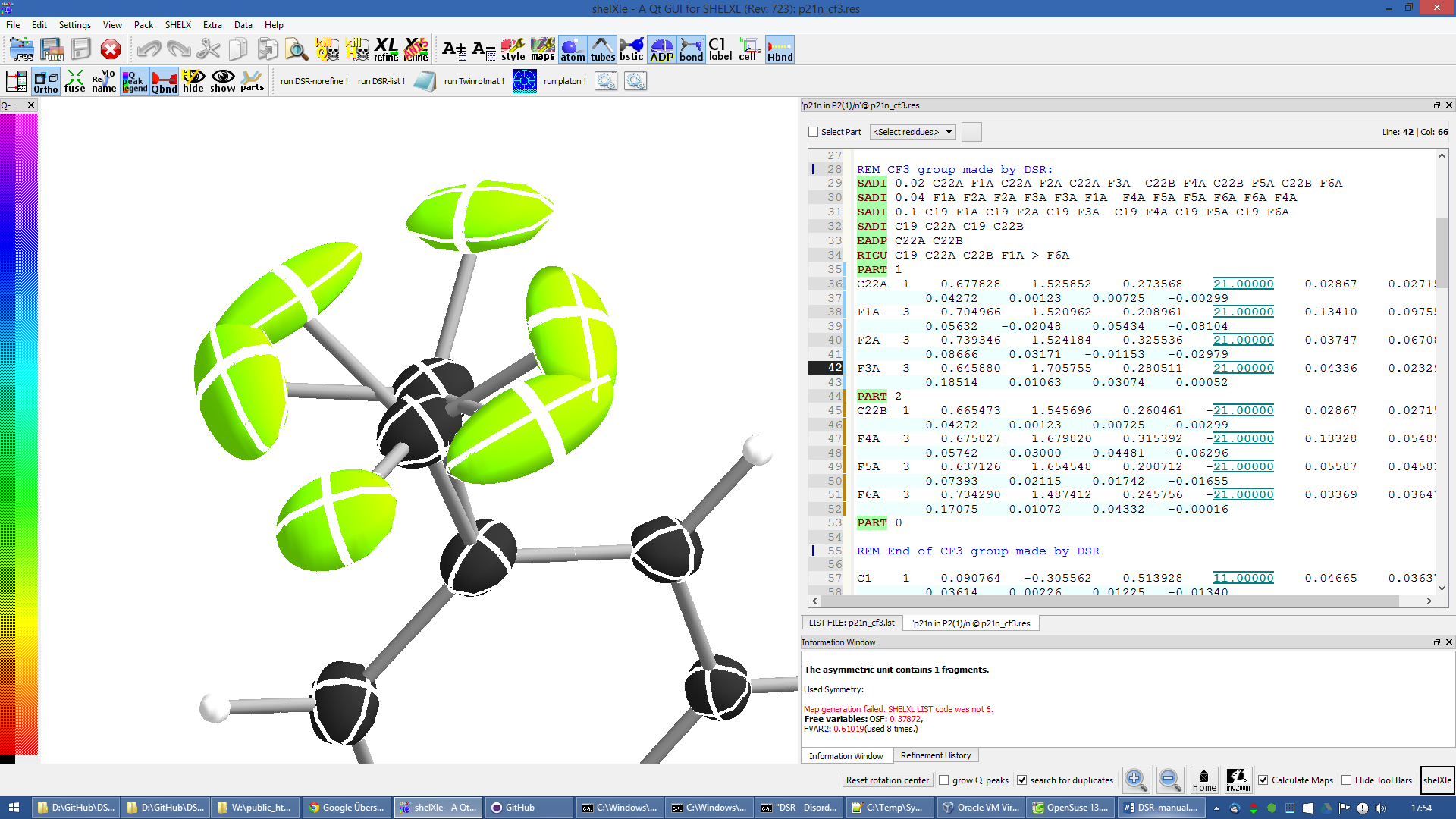
Figure 14 CF3 group split on two positions (rem dsr put CF6 on C22 split).
You should never use CF9 just because it is possible! Often CF6 is sufficient. CF9 often just uses more least-squares parameters without improving the model.
You can try the above example with the structure “p21n_cf3.res” located in the example directory of DSR. At the end of the file, there is already a command line to place a CF3 group on C22.
Tips and Tricks
You can quickly rename molecular moieties using DSR in the replace mode. This is useful directly after the solution with SHELXT for example. In the replace mode, DSR replaces all atoms that are in PART 0 and that are 1.3 Å near the atoms of the placed fragment. Using replace mode prevents you from renaming every single atom.
You want to know typical bond lengths of fragments included in the DSR database? Export the respective fragment and either see 1,2 and 1,3 distances in the restraints lists or let the SHELX GUI of your choice find out any distance of the respective atoms.
The command “dsr –l” will tell you about errors in the fragment database in case you created a fragment yourself and made a mistake.
The ShelXle interface allows you to edit or create fragments. In the rename mode, you can rename atoms of a fragment and their respective restraints.
In ShelXle, it is not necessary to do anything special for atoms on special positions anymore. Every symmetry related atom or q-peak (grow q-peaks option) can be chosen as target position for DSR. DSR uses the coordinates instead of atoms names.
In ShelXle, you can easily go back in the DSR model with the “restore last .res” button. This restores the .res file from just before the fragment fit. The complete save history in ShelXle is in: menu→file→open save history.
Fragments Included in the Database
Type “dsr –l” to see a list of fragments supplied with DSR. Also the graphical user interfaces show a list of all fragments.
For Developers
Graphical user interfaces for SHELXL (like ShelXle) can use several special commands to control DSR:
$ dsr –lc
DSR version: 199
acetate;;Acetate anion, C2H3O2-;;1105;;dsr_db
acetone;;Acetone, C3H6O;;2312;;dsr_db
acetonitrile;;Acetonitrile, C2H3N, NMe;;1670;;dsr_db
adamantane;;Adamantane, C10H16;;2405;;dsr_db
...
The first line of the –lc output is the version of DSR. All other lines are separated by double semicolon. Each fragment uses one line. The first column is the name tag, the second is the full name and the third is the type of database (dsr_db/dsr_usr_db).
$ dsr -x tol
toluene;;Toluene, C7H8;;885;;dsr_db
tbu-o;;tert-Butanol (or tert-Butoxy);;2203;;dsr_db
tosylate;;Tosylate anion, CH3C6H4SO3-;;2447;;dsr_db
dmpz;;Dimethylpyrazolato anion, [C5H6N]-, pz*;;1126;;dsr_db
ile;;ISOLEUCINE;;4327;;dsr_db
pyr;;PYRROLYSINE;;3802;;dsr_db
acetonitrile;;Acetonitrile, C2H3N, NMe;;1670;;dsr_db
mquinolinol;;2-Methyl-8-quinolinol, C10H9NO;;2041;;dsr_db
The –x parameter searches for fragments and displays the result with the same syntax as –lc.
$ dsr -ah toluene
<atoms>
C1 6 1.78099 7.14907 12.00423;;C2 6 2.20089 8.30676 11.13758;;C3 6
1.26895 9.02168 10.39032;;C4 6 1.64225 10.07768 9.58845;;C5 6 2.98081
10.44432 9.51725;;C6 6 3.92045 9.74974 10.25408;;C7 6 3.53891 8.69091
11.05301
</atoms>
<tag>
toluene
</tag>
<comment>
Toluene, C7H8
</comment>
<source>
CCDC CESLUJ
</source>
<cell>
1;;1;;1;;90;;90;;90
</cell>
<residue>
TOL
</residue>
<dbtype>
dsr_db
</dbtype>
<restr>
SADI C2 C3 C3 C4 C4 C5 C5 C6 C6 C7 C7 C2;;SADI 0.04 C2 C6 C2 C4 C7 C5 C3
C7 C4 C6 C3 C5;;DFIX 1.51 C1 C2;;SADI 0.04 C1 C7 C1 C3;;FLAT C1 > C7;;
SIMU C1 > C7;;RIGU C1 > C7
</restr>
The –ah command displays the content of the database entry for the respective fragment on screen. Each type of database entry has a start end an end tag. Each entry is a single delimited by double semicolon.
-shx "c:\Program files\shelx\shelxl"
Command to tell DSR where SHELXL is located.
$ dsr –ea
Exports all fragments at once to .res files in the current directory.
$ dsr –target [coordinate triples]
With the target option you are able to define the exact coordinates of the target position for the fragment fit. Coordinates are given in space separated coordinates.
General Remarks
Parsers for DSR output should be aware that DSR prints error messages between three stars (SHELXL uses two stars). For example:
*** Check database entry. ***